February 17, 2016
RNA Modification Discovery Suggests New Code for Control of Gene Expression
Congratulations to Dan Dominissini of The University of Chicago! Dominissini is the first author on a recent report published in Nature: “The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA.” He is a postdoctoral fellow in Chuan He’s laboratory and recipient of a 2014 CBC Postdoctoral Research Award. The report is a collaborative effort by scientists from Tel Aviv University, Sheba Medical Center, and the University of Chicago. The authors describe a new mRNA modification, N1-methyladenosine (m1A), that is evolutionarily conserved from yeast to humans, is enriched around start codons and appears to promote mRNA translation.
A new cellular signal discovered by a team of scientists at the University of Chicago and Tel Aviv University provides a promising new lever in the control of gene expression.
The study, published online Feb. 10 in the journal Nature, describes a small chemical modification that can significantly boost the conversion of genes to proteins. Together with other recent findings, the discovery enriches a critical new dimension to the “Central Dogma” of molecular biology: the epitranscriptome.
 “This discovery further opens the window on a whole new world of biology for us to explore,” said Chuan He, the John T. Wilson Distinguished Service Professor in Chemistry, Howard Hughes Medical Institute investigator and senior author of the study. “These modifications have a major impact on almost every biological process.” Dan Dominissini (right), a postdoctoral fellow in Chuan He’s group, is the first author of the publication.
“This discovery further opens the window on a whole new world of biology for us to explore,” said Chuan He, the John T. Wilson Distinguished Service Professor in Chemistry, Howard Hughes Medical Institute investigator and senior author of the study. “These modifications have a major impact on almost every biological process.” Dan Dominissini (right), a postdoctoral fellow in Chuan He’s group, is the first author of the publication.
The central dogma of molecular biology describes the cellular pathway where genetic information from DNA is copied into temporary RNA “transcripts,” which provide the recipe for the production of proteins. Since Francis Crick first postulated the theory in 1956, scientists have discovered a multitude of modifications to DNA and proteins that regulate this process.
Only recently, however, have scientists focused on investigating dynamic modifications that specifically target the RNA step. In 2011, He’s group discovered the first RNA demethylase that reverses the most prevalent mRNA methylation N6-methyladenosine, or m6A, implying that the addition and removal of the methyl group could dramatically affect these messengers and impact the outcome of gene expression, as also seen for DNA and histones. Subsequently, scientists discovered that the dynamic and reversible methylation of m6A dramatically controlled the metabolism and function of most cellular messenger RNA, and thus, the production of proteins.
 In the new Nature study, researchers from UChicago and Tel Aviv University describe a second functional mRNA methylation, N1-methyladenosine, or m1A. Like m6A, the small modification is evolutionarily conserved and common, and present in humans, rodents and yeast, the authors found. But its location and effect on gene expression reflect a new form of epitranscriptome control and suggest an even larger cellular “control panel.”
In the new Nature study, researchers from UChicago and Tel Aviv University describe a second functional mRNA methylation, N1-methyladenosine, or m1A. Like m6A, the small modification is evolutionarily conserved and common, and present in humans, rodents and yeast, the authors found. But its location and effect on gene expression reflect a new form of epitranscriptome control and suggest an even larger cellular “control panel.”
“The discovery of m1A is extremely important, not only because of its own potential in affecting biological processes, but also because it validates the hypothesis that there is not just one functional modification,” He said. “There could be multiple modifications at different sites where each may carry a distinct message to control the fate and function of mRNA.”
The researchers estimated that m1A was present on transcripts of more than one out of three expressed human genes. Methylated genes exhibited enhanced translation compared to unmethlyated genes, producing protein levels nearly twice as high in all cell types. This increase suggests that m1A, like m6A, may be a mechanism by which cells rapidly boost the expression of hundreds or thousands of specific genes, perhaps during important processes such as cell division, differentiation or under stress.
“mRNA is the perfect place to regulate gene expression, because they can code information from transcription and directly impact translation; you can add a consensus sequence to a group of genes and use a modification of the sequence to readily control several hundred transcripts simultaneously,” He said. “If you want to rapidly change the expression of several hundred or a thousand genes, this offers the best way.”
However, despite their complementary effects, m1A and m6A exert their influence on mRNA through different pathways. While studies have found that m6A localizes predominantly to the tail of messenger RNA molecules, increasing their translation and turnover rate, m1A was found largely near the start codon of mRNA transcripts, where protein translation begins. The different mechanisms could allow for finer tuning of post-transcriptional gene expression, or the selective activation of particular genes in different physiological situations.
“This study represents a breakthrough discovery in the exciting, nascent field of the ‘epitranscriptome,’ which is how RNAs are regulated, akin to the genome and the epigenome,” said Christopher Mason, associate professor at Weill Cornell Medicine, who was not affiliated with the study. “What is important about this work is that m6A was recently found to enrich at the ends of genes, and now we know that m1A is what is helping regulate the beginning of genes, and this opens up many questions about revealing the ‘epitranscriptome code’ just like the histone code or the genetic code.”
Future studies will examine the role of m1A methylation in human development, diseases such as diabetes and cancer, and its potential as a target for therapeutic uses.
The study was supported by the Kahn Family Foundation, the Sagol Neuroscience Network, the National Institutes of Health, Howard Hughes Medical Institute, Flight Attendant Medical Research Institute, Israel Science Foundation, Israeli Centers of Excellence Program, Ernest and Bonnie Beutler Research Program, Chicago Biomedical Consortium, and the Damon Runyon Cancer Research Foundation.
Adapted (with modifications) from: UChicago News, by Robert Mitchum, February 16, 2016
Photo: Dan Dominissini, Courtesy of Science 05 Dec 2014: Vol. 346, Issue 6214, pp. 1192
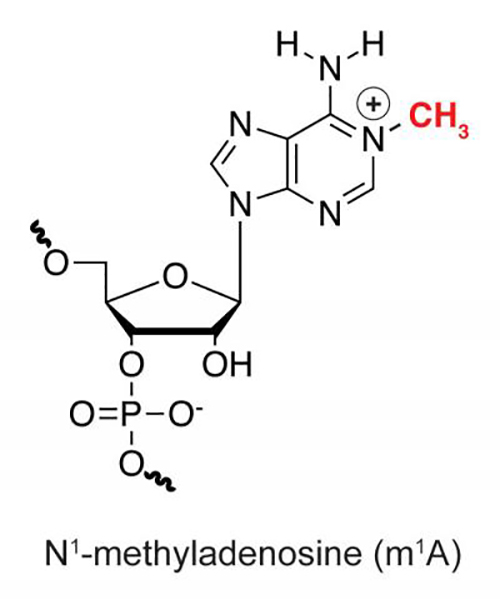
Figure: The discovery of a new messenger RNA (mRNA) modification m1A and its distinct location in mammalian mRNA, together with previously known mRNA modifications, strongly suggest the presence of "epitranscriptome code" in controlling gene expression. This diagram of an RNA molecule shows the spot where the CH3 methyl group is added for an N1 methylation, which adds a positive charge. Courtesy of Chuan He/University of Chicago
CBC Postdoctoral Award (2014): Dan Dominissini, PI: Chuan He, University of Chicago, for project: Pseudouridine Sequencing – a Roadmap to the New Biology of mRNA Pseudouridine Modification
Publication attributed to the CBC funding: Dominissini D, Nachtergaele S, Moshitch-Moshkovitz S, Peer E, Kol N, Ben-Haim MS, Dai Q, Di Segni A, Salmon-Divon M, Clark WC, Zheng G, Pan T, Solomon O, Eyal E, Hershkovitz V, Han D, Doré LC, Amariglio N, Rechavi G, He C. The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA. Nature. 2016 Feb 10. [Epub ahead of print] (PubMed)
